What is gene ontology?
Gene ontology considers the interaction of large functional networks of gene products. Databases such as The Gene Ontology create networks of gene product annotations that can be searched to find information about development within a variety of different model organisms. The functional annotations within the Gene Ontology involve three components: molecular function (which describes the biochemical function of a gene product carried out on a molecular level), biological process (which describes the biological objective of a gene product), and cellular localization (which describes where in a cell a gene product localizes) [2].
Molecular Function of polycystin-1
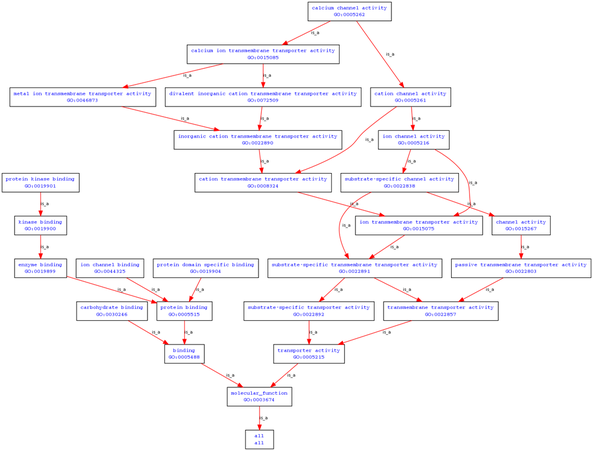
Examination of polycystin-1's molecular function via the Gene Ontology indicates that polycystin-1 functions in calcium channel activity, carbohydrate binding, protein binding, and protein kinase binding [3].
These annotations make sense, given the variety of extracellular domains present on polycystin-1. The extracellular domains on polycystin-1 have been implicated in carbohydrate binding, and polycystin-1 is expected to interact with polycystin-2 in calcium channel activity.
These annotations make sense, given the variety of extracellular domains present on polycystin-1. The extracellular domains on polycystin-1 have been implicated in carbohydrate binding, and polycystin-1 is expected to interact with polycystin-2 in calcium channel activity.
Cellular localization of polycystin-1
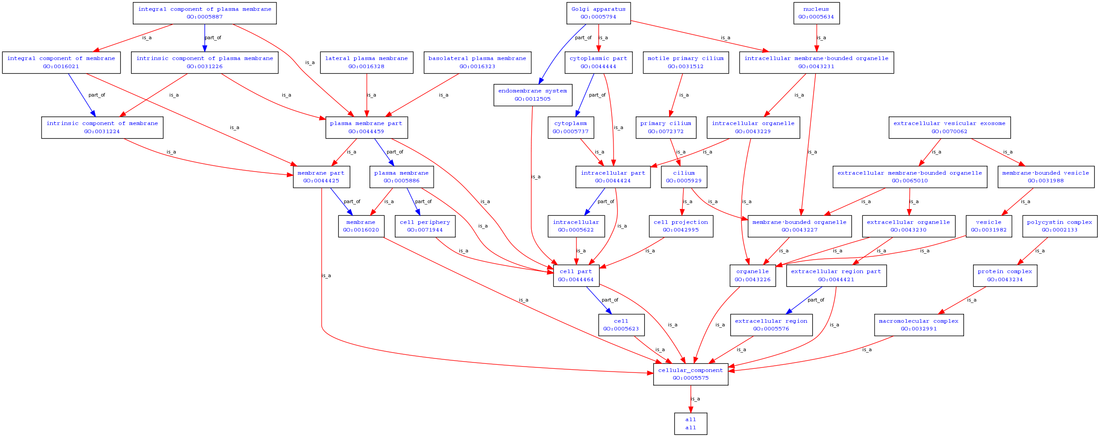
Examination of polycystin-1's cellular function via the gene ontology indicate that polycystin-1 has been found in the basolateral plasma membrane, cilium, cytoplasm, the extracellular vesicle exosome, the lateral plasma membrane, primary cilium, the golgi apparatus, and the nucleus.
These connections generally make sense, given that polycystin-1 is primarily localized to the membrane of primary cilia and that it can undergo proteolytic cleavage whereby the c-terminal tail translocates to the nucleus and initiates signalling processes [4].
These connections generally make sense, given that polycystin-1 is primarily localized to the membrane of primary cilia and that it can undergo proteolytic cleavage whereby the c-terminal tail translocates to the nucleus and initiates signalling processes [4].
Biological function of polycystin-1
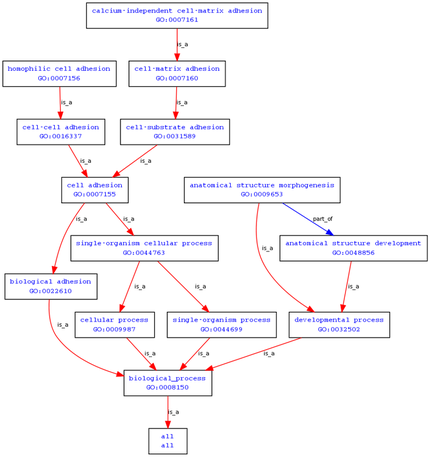
Querying the database for biological process associations yields a number of associations, although many of them are predicated and not presently validated by research. The terms that have been validated by research include: anatomical structure morphogenesis, cell-matrix adhesion, and homophilic cell adhesion.
These associations are interesting, although they offer minimal information about the significance of polcystin-1 in the development of polycystic kidney disease and its ultimate impact on signaling.
These associations are interesting, although they offer minimal information about the significance of polcystin-1 in the development of polycystic kidney disease and its ultimate impact on signaling.
Analysis
Gene Ontology analysis of PKD1 yields a large number of results, likely due to the size of the gene, and the prevalence of the mutations. These annotations generally make sense and support the role of polycystin-1 in signaling within primary cilia. Interestingly, the biological function annotations of PKD1 are less thorough and comprehensive than the molecular function and cellular component annotations, which probably reflects current uncertainty regarding the role that polycystin-1 plays in normal development and the aberrant growth witnessed upon PKD1 mutation.
References[1] Banner image: http://www.biomedcentral.com/1752-0509/2/25/figure/F1?highres=y
[2] Hill, D., Berardini, T., Howe, D., Van Auken, K. (2010). Representing ontogeny through ontology: A developmental biologist's guide to the gene ontology. Molecular reproduction and development (77) 314-329. [3] PKD1: The Gene ontology [4] Chauvet, V., Tian, X., Husson, H., Grimm, DH., Wang, T., Hiesberger, T., Igarashi, P., Bennett, AM., Ibraghimov-Beskrovnaya, O., Somlo, S., Caplan, MJ. (2005) Mechanical stimuli induce cleavage and nuclear translocation of the polycystin-1 c-terminus. J clin invest. 114(10) 1433-43. |
Site created by: Elizabeth Roeske Last updated: 5.12.2014 University of Wisconsin-Madison: Genetics 564 |