This web page was produced for Genetics 564, an undergraduate course at UW-Madison.
What is Homology?
Homology can be conceptualized as similarity resulting from common ancestry. The remarkable similarity of organisms (even between those organisms that are distantly related) provides evidence for evolution while also yielding the opportunity to make testable predictions about function or explain observations that might otherwise be puzzling [2]. To examine the similarity between PDK1 proteins in different organisms , the nucleotide sequences were aligned using NCBI's basic alignment search tool (BLAST). BLAST compares a query sequence (in this case, human PDK1) to a set of reference genomes within other organisms in order to find similar sequences. The program is also able to provide a measure of the statistical significance of relationships between sequences. In order to assure high confidence of homologues, Homologene was used to confirm likely homologues initially identified using BLAST.
Analysis of PKD1 gene homology
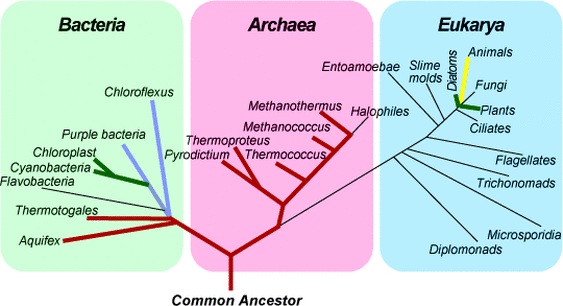
The PKD1 gene is conserved across species, as measured by comparative sequence analysis, although the level of this conservation varies usually based on the organism's evolutionary relationship to humans. PKD1 can be found in Eukarya, although not in Bacteria or Archaea. The gene is especially conserved in Euteleostomi, which is clade that includes 90% of living vertebrates [3]. These results generally make sense, because PKD1 is found within primary (non-motile) cilia, which are not found in bacteria and are found only rarely in plants.
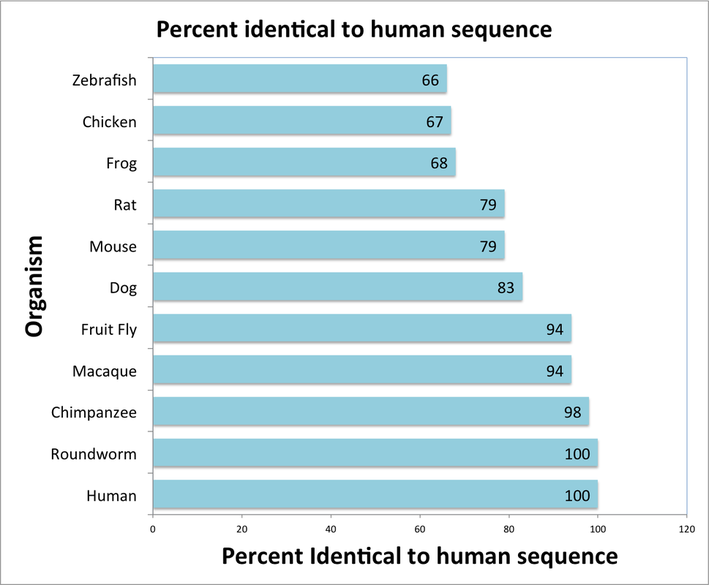
Comparison of each species' similarity to the human reference sequence. More closely evolutionarily related versions of PKD1 share more sequence similarity.
Animals
References
|
Site created by: Elizabeth Roeske Last updated: 5.12.2014 University of Wisconsin-Madison: Genetics 564 |