What are protein networks?
Protein networks describe the organization that allows proteins to work cooperatively to form molecular machines and the functional networks that control cellular behavior. Protein-protein interactions can be identified in a number of different ways. Yeast two-hybrid screens allow for the detection of binary interactions between two proteins. Visualizing protein networks may help to examine biochemical cascades or better understand disease pathogenesis.
|
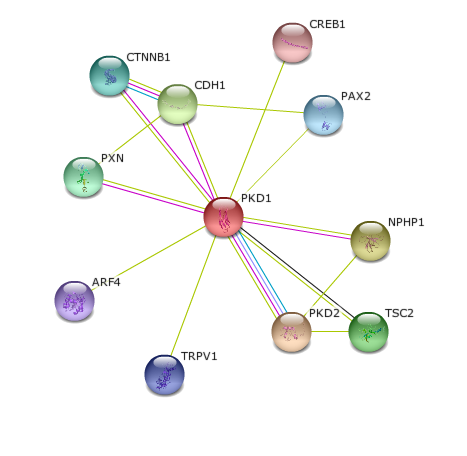
Polycystin-1 has several identifiable interaction partners in humans:
CREB1: This protein binds the cAMP response element (CRE), a sequence present in many viral and cellular promoters. CREB stimulates transcription on binding to the CRE. PAX2: Probable transcription factor that may have a role in kidney cell differentiation. Has a critical role in the development of the urogenital tract, the eyes, and the CNS NPHP1: Together with Cas it may play a role in the control of epithelial cell polarity. PKD2: Functions as a calcium permeable cation channel. PKD1 and PKD2 may function through a common signaling pathway that is necessary for normal tubulogenesis TSC2: Implicated as a tumor suppressor. Involved in microtubule-mediated protein transport TRPV1: Receptor-activated non-selective calcium permeant cation channel involved in detection of noxious chemical and thermal stimuli. ARF4: Involved in protein trafficking; may modulate vesicle budding and uncoating within the Golgi apparatus PXN: Cytoskeletal protein involved in actin-membrane attachment at sites of cell adhesion to the extracellular matrix (focal adhesion) CTNNB1: Involved in the regulation of cell adhesion and in signal transduction through the Wnt pathway CDH!: CDH1 is involved in mechanisms regulating cell-cell adhesions, mobility and proliferation of epithelial cells. |
Analysis of this network
Many of these interaction partners are consistent with terms previously identified using gene ontology, which reinforces the role of PKD1 in a pathway involving extracellular sensation and signaling within the cell. For example, gene ontology terms implicated cation channels and signaling. Interestingly, PAX2 provides a possible link to early development, although other aging and development proteins were not identified through this analysis of gene interaction. Additionally, as seen here, there is a robust association between PKD1 and PKD2, which is consistent with extensive literature detailing the close interplay between PKD1 and PKD2.
|
References
[1] Banner image: http://gmdd.shgmo.org/Computational-Biology/ANAP/ANAP_V1.1/help/anap-userguide/manual.html |
Site created by: Elizabeth Roeske
Last updated: 5.12.2014 University of Wisconsin-Madison: Genetics 564 |